Description: Participants: Interactions 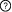 This is a sequential switch. The interactions in this switch take place in a defined order. The order is defined by numbers to the left of each interaction. More than one interaction can occur in each step, these interactions occur independently but are necessary for future step This is a sequential switch. The interactions in this switch take place in a defined order. The order is defined by numbers to the left of each interaction. More than one interaction can occur in each step, these interactions occur independently but are necessary for future step1Interaction #1 YWHAB - YWHAB Interfaces (1) 14-3-3 protein in 14-3-3 protein beta/alpha (YWHAB) (2) 14-3-3 protein (5-238) in 14-3-3 protein beta/alpha (YWHAB) 2Interaction #2 FOXO3 - Interaction #1 YWHAB - YWHAB Requires Interaction #1 YWHAB - YWHAB Interfaces (3) LIG_14-3-3_3 motif (250RAVSMD255) in Forkhead box protein O3 (FOXO3) (4) Interaction #1 YWHAB - YWHAB Interaction Regulation PTM-dependent Induction (Phosphorylation of S253 on Forkhead box protein O3 (FOXO3)) of the Forkhead box protein O3 (FOXO3) LIG_14-3-3_3 motif - 14-3-3 protein interaction Regulatory Enzymes for switch Modifying enzymes for residue: S253: RAC-alpha serine/threonine-protein kinase (AKT1) 2Interaction #3 FOXO3 - Interaction #1 YWHAB - YWHAB Requires Interaction #1 YWHAB - YWHAB Interfaces (5) LIG_14-3-3_3 motif (29RSCTWP34) in Forkhead box protein O3 (FOXO3) (6) Interaction #1 YWHAB - YWHAB Interaction Regulation PTM-dependent Induction (Phosphorylation of T32 on Forkhead box protein O3 (FOXO3)) of the Forkhead box protein O3 (FOXO3) LIG_14-3-3_3 motif - 14-3-3 protein interaction Regulatory Enzymes for switch Modifying enzymes for residue: T32: RAC-alpha serine/threonine-protein kinase (AKT1) Inferred Regulatory Enzymes for switch Putative modifying enzymes for residue: T32 : PKB_group, SGK_group, RAC-alpha serine/threonine-protein kinase (AKT1), RAC-alpha serine/threonine-protein kinase (Akt1), Serine/threonine-protein kinase pim-1 (PIM1). Context Switch localisation ELM curated: nucleus, cytosol, internal side of plasma membrane Switch pathways Curated: PI3K-Akt signaling pathway References (1) A conserved MST-FOXO signaling pathway mediates oxidative-stress responses and extends life span. See also Other switches involving participants |
|
Please send any suggestions/comments to: switches@elm.eu.org

